You have no items in your shopping cart.
CUT&RUN vs CUT&Tag
Technical Comparison, Key Applications, and How to Choose the Right Chromatin Profiling Method
Overview: CUT&RUN and CUT&Tag have become two of the most important next-generation alternatives to traditional ChIP-seq for profiling protein–DNA interactions and chromatin states. Both methods offer significantly lower input requirements, improved signal-to-noise ratios, and higher resolution than conventional ChIP-seq workflows.
Although the two techniques are closely related in principle, they differ in the enzyme used, the way DNA is processed, the signal pattern produced, and the experimental questions they are best suited to answer.
In this guide, we compare CUT&RUN, CUT&Tag, and traditional ChIP-seq, outline their major strengths and limitations, and provide a practical framework for selecting the most suitable method for your research.
What are CUT&RUN and CUT&Tag?
CUT&RUN (Cleavage Under Targets & Release Using Nuclease) and CUT&Tag (Cleavage Under Targets & Tagmentation) are antibody-guided chromatin profiling methods designed to map histone modifications, transcription factor binding, and chromatin-associated proteins with high precision.
In both methods, a target-specific antibody first binds the chromatin-associated protein of interest in situ. A fusion enzyme is then recruited to the antibody complex, where it acts locally to generate DNA fragments suitable for downstream sequencing.
The central technical distinction lies in the effector enzyme. CUT&RUN uses a protein A/G-MNase fusion protein, while CUT&Tag uses a protein A/G-Tn5 transposase fusion. This difference strongly influences fragment generation, library preparation, signal shape, and ultimately the biological interpretation of the data.
How does CUT&RUN work?
CUT&RUN employs a protein A/G-linked micrococcal nuclease (pA/G-MNase). For researchers seeking highly reproducible fragmentation, workflows frequently utilize optimized master mixes like the Vazyme Hyperactive pG-MNase CUT&RUN Assay Kit for Illumina.
Once guided to the antibody-bound target, the MNase is activated by calcium and cleaves DNA in the immediate vicinity of the binding event. The released DNA fragments diffuse into the supernatant and are then collected for standard library preparation. Because MNase remains enzymatically active during the reaction, CUT&RUN often generates a broader range of DNA fragment sizes and a more extended signal profile across chromatin regions.
Mild formaldehyde crosslinking is commonly used to preserve nuclear integrity during the workflow, although the procedure remains substantially gentler than traditional ChIP-seq.
How does CUT&Tag work?
CUT&Tag uses a protein A/G-linked Tn5 transposase (pA/G-Tn5). To ensure maximal tagmentation efficiency and minimal background, many laboratories rely on specialized, all-in-one systems such as the Vazyme Hyperactive Universal CUT&Tag Assay Kit for Illumina Pro.
After antibody-guided recruitment to the chromatin target, the enzyme is activated by magnesium. It simultaneously cleaves the DNA and inserts sequencing adapters directly into the target-associated genomic region. Because Tn5 performs a highly localized insertion event, CUT&Tag typically produces sharper peaks and lower background than CUT&RUN.
Another major practical advantage is that adapter insertion occurs during the assay itself, simplifying downstream library preparation and making the workflow particularly attractive for low-input and single-cell applications. CUT&Tag is usually performed under native, non-crosslinked conditions, helping preserve native chromatin conformation.
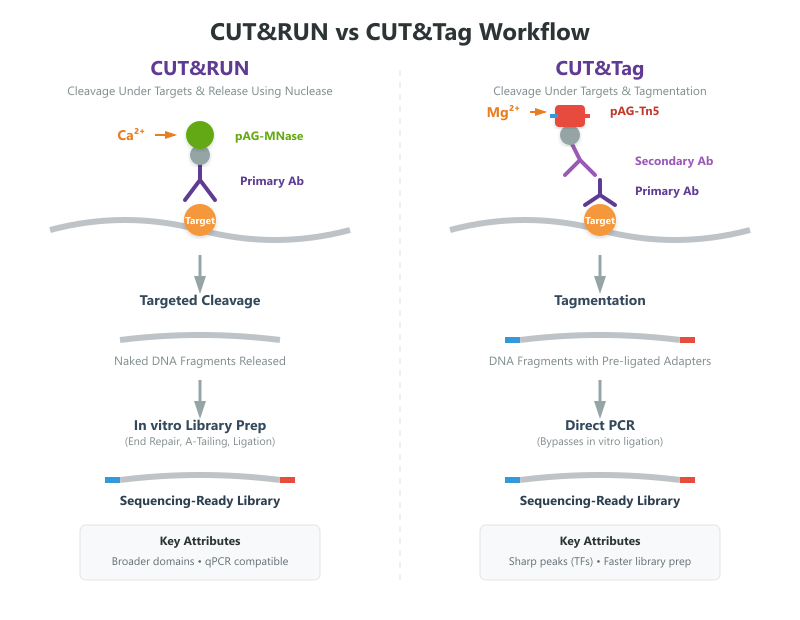 CUT&RUN vs CUT&Tag
CUT&RUN vs CUT&Tag
CUT&RUN vs CUT&Tag vs ChIP-seq: Key Technical Differences
Traditional ChIP-seq has long been the standard method for profiling chromatin-associated factors, but it generally requires far larger sample input, relies on sonication-based fragmentation, and often produces higher background with lower spatial resolution.
- CUT&RUN uses pA/G-MNase, mild crosslinking, and standard library preparation. It typically works well with approximately 50,000–300,000 cells and produces high signal-to-noise with relatively broad signal profiles.
- CUT&Tag uses pA/G-Tn5, native conditions, and integrated tagmentation-based library preparation. It is compatible with much lower input, often around 5,000–50,000 cells, and can be adapted for single-cell workflows.
- ChIP-seq generally requires millions of cells, formaldehyde crosslinking, sonication, and a longer workflow, while often delivering more moderate resolution and signal quality.
From a practical perspective, both CUT&RUN and CUT&Tag can usually be completed within 1–1.5 days, whereas traditional ChIP-seq workflows often extend to 2–3 days.
Signal Characteristics: Broad Domains vs Sharp Peaks
One of the most important differences between CUT&RUN and CUT&Tag is the shape of the signal they generate across the genome.
For broad histone modifications such as H3K27me3, CUT&RUN typically produces a more extended, plateau-like distribution that better reflects large regulatory or repressive chromatin domains. This makes it especially useful when the biological question concerns the overall size, spread, or architecture of a domain rather than a single sharp binding event.
CUT&Tag, by contrast, tends to concentrate signal tightly around the center of the target region. This produces sharper peaks with weaker peripheral signal, which can be extremely advantageous when precise localization is required.
For narrow binding proteins such as transcription factors or chromatin regulators with focused occupancy patterns, CUT&Tag typically delivers a clear resolution advantage. In these settings, it can localize binding sites with very high precision, often within tens of base pairs, whereas CUT&RUN signals are somewhat broader.
Key Research Tools & Kits for Chromatin Profiling
The performance of CUT&RUN and CUT&Tag depends heavily on reagent quality, especially antibody specificity, enzyme efficiency, and lot-to-lot consistency.
For laboratories looking to standardize their epigenomic assays, utilizing all-in-one solutions like the Vazyme CUT&Tag Assay Kit or the Vazyme CUT&RUN Assay Kit ensures high-fidelity sequencing libraries with minimal optimization. At Biorbyt, these kits perfectly complement our broad range of primary antibodies tailored for epigenetic target analysis.
Quantitative Performance and Data Quality
Quantitative comparisons further highlight the practical differences between the two methods. CUT&Tag commonly produces higher FRiP (Fraction of Reads in Peaks) values, indicating a greater proportion of reads falling within enriched regions, and generally exhibits lower background noise.
CUT&RUN, however, often shows broader peak widths, which is consistent with its strength in capturing extended chromatin domains rather than only sharp signal maxima.
In simple terms, CUT&Tag often provides maximum precision and sensitivity, while CUT&RUN offers stronger representation of broad chromatin context.
When should you choose CUT&RUN?
CUT&RUN is often the preferred method when the research goal is to investigate broad chromatin regulatory landscapes rather than pinpointing an exact nucleotide-scale binding event.
- Profiling broad histone modifications such as H3K27me3 or H3K9me3
- Studying repressive chromatin domains and their expansion or contraction during development or disease
- Investigating intergenic regulatory regions or extended domain-level chromatin organization
- Generating high-quality chromatin profiles from lower input than ChIP-seq, while retaining strong domain-level context
Researchers interested in epigenetic state changes across large genomic territories often find CUT&RUN especially informative.
When should you choose CUT&Tag?
CUT&Tag is typically the better option when highly focused signal, minimal background, and maximal sensitivity are the priority.
- Mapping transcription factor binding sites with high positional precision
- Profiling promoter-associated histone modifications such as H3K4me3
- Working with extremely limited material, including rare primary cells or single-cell applications
- Reducing library preparation complexity through direct adapter insertion
If the key question is, “Where exactly does this factor bind?”, CUT&Tag is often the strongest choice.
Typical Applications
Both methods are widely used to detect chromatin-associated biological events with far greater efficiency than many conventional immunoprecipitation-based workflows.
- Histone modification profiling, including active and repressive chromatin marks
- Transcription factor binding analysis
- Mapping chromatin-associated proteins such as CTCF, EZH2, and RNA polymerase
- Profiling rare cell populations or low-input clinical samples
- Early developmental or embryonic chromatin studies
- Single-cell epigenomics and high-resolution chromatin state analysis
Practical Decision Guide
A useful way to think about these methods is to match the technique to the biological question.
- Choose CUT&RUN when the question is about broad chromatin architecture, domain-level regulatory states, or spreading histone marks.
- Choose CUT&Tag when the question is about highly precise localization, narrow peaks, or the most efficient use of very limited input material.
- Choose ChIP-seq when legacy comparability, established historical datasets, or validated existing pipelines are the primary consideration.
In many projects, the two methods should be viewed as complementary rather than competing. CUT&RUN may provide better large-scale chromatin context, while CUT&Tag may offer sharper target localization for focused mechanistic studies.
References
- Oh, Y., et al. Comparative analyses of ChIP-seq, CUT&RUN and CUT&Tag for Polycomb chromatin profiling. BMB reports, p.6727.
- Tao X., et al. (2020). Efficient chromatin profiling of H3K4me3 modification in cotton using CUT&Tag. Plant Methods. 16:120. doi: 10.1186/s13007-020-00664-8. PMID: 32884577.
- Ito S., et al. (2024). Polycomb-mediated histone modifications and gene regulation. Biochem Soc Trans. 52(1):151-161. doi: 10.1042/BST20230336. PMID: 38288743.
