You have no items in your shopping cart.
10X PBS pH 7.2
Description
Images & Validation
−| Application Notes |
|---|
Key Properties
−| Purity | 10X PBS buffer was aseptically filtered through a Millipore 0.22 micron filter into clean, pre-sterilized containers. The product was tested on trypticase soy agar for 24 hours, 48 hours and 72 hours and was found to be negative for bacteria. |
|---|---|
| Conjugation | Unconjugated |
Storage & Handling
−| Storage | Store container at room temperature (18° to 26° C) prior to opening. If desired, the solution may be stored at 4° C or less. Some salts may precipitate out of solution at lower temperature. Allow buffer to equilibrate to room temperature (18° to 26° C) to restore solubility of some salts. |
|---|---|
| Form/Appearance | Liquid |
| Buffer/Preservatives | Preservative: None. Stabilizer: None; Buffer: See application note. |
| Concentration | 10X |
| Expiration Date | 12 months from date of receipt. |
| Hazard Information | Non-Toxic |
| Disclaimer | For research use only |
Alternative Names
−Similar Products
−
Quality Guarantee
Explore bioreagents carefree to elevate your research. All our products are rigorously tested for performance. If a product does not perform as described on its datasheet, our scientific support team will provide expert troubleshooting, a prompt replacement, or a refund. For full details, please see our Terms & Conditions and Buying Guide. Contact us at [email protected].
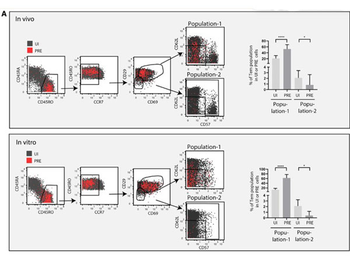
A subset of Tem-like cells sorted based on surface markers defining clusters 12 and 13 are highly susceptible to HIV infection. (A) Shown are the CyTOF datasets, with UI CD4+ T cells shown in gray and the HIV-susceptible PRE cells shown in red. Cells were pre-gated on live, singlet CD3 + CD19−CD8−CD4+ T cells. A sequential gating strategy was then implemented using surface markers characteristic of HIV-susceptible cells as defined by clusters 12 and 13. This strategy was used to characterize a final population of "population-1" cells (CD3 + CD4+ CD45RO + CD45RA−CCR7low/medCD29med/highCD69med/high CD62LlowCD57low/med), which were more abundant among PRE cells than among UI cells. For comparison, we characterized a "population-2" (CD3 + CD4+ CD45RO + CD45RA−CCR7low/medCD29lowCD69low and not CD62LlowCD57low/med) predicted to be much less susceptible to infection because it comprised a significantly lower proportion of PRE cells. The gating strategies are shown on the left, whereas the graphs on the right depict the frequencies of the population-1 and population-2 subsets within the UI and PRE cell populations. Note that the over-representation of population-1 cells among PRE cells suggest their preferential susceptibility to infection, whereas the under-representation of population-2 cells among PRE cells suggest their relative resistance to infection. *p < 0.05, ****p < 0.0001 as determined by a Student's paired t test. Error bars correspond to the standard deviation.
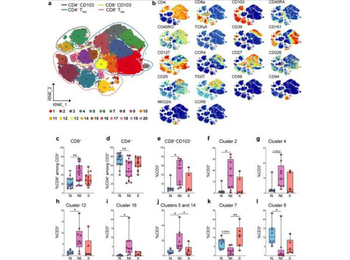
Quantification of CD8 and CD4 T cell clusters by CyTOF analysis in LP of control and CD patients. a Schematic t-SNE of CD4+ and CD8+ T cells from LP of all donors concatenated together (n = 18) controls (N), n = 8; CD, non-inflamed site (NI), n = 9; CD, inflamed site (II), n = 6. Total of 23 samples. b t-SNE of the indicated markers in CD4+ and CD8+ T cells. c, d Quantification of total CD8+ (c), total CD4+ (d) in LP of controls and CD patients by CyTOF (triangles = fresh samples) and FACS (circles = frozen samples). c, d Control (N), n = 17 (8 fresh, 9 frozen); CD, non-inflamed site (NI) n = 19 (9 fresh, 10 frozen); CD, inflamed site (II), n = 14 (6 fresh, 8 frozen). e–i Quantification of total CD8+ TRM (e) and CD8+ clusters 2 (f), 4 (g), 12 (h), and 16 (i) in LP of controls and CD patients by CyTOF. j–l Quantification of the CD4+ clusters 5 and 14 (j), 7 (k), and 9 (l) in LP of controls and CD patients by CyTOF. e–l Controls (N), n = 8; CD, non-inflamed site (NI), n = 9; CD, inflamed site (II), n = 6. Circles and triangles on the boxplots show data collected for each individual donor. Data were median and interquartile range. Significance was calculated using an ordinary, one-way ANOVA, multiple comparisons test with Prism v8 software. c **P = 0.0014; d **P = 0.028; e *P = 0.0139; f *P = 0.0178; h *P = 0.0178; i *P = 0.0219; j N vs. NI *P = 0.0156, NI vs. II *P = 0.0465; k **P = 0.0014; l **P = 0.0283. TRM tissue-resident memory T cell.
Documents Download
Request a Document
Protocol Information
10X PBS pH 7.2 (orb348532)
Participating in our Biorbyt product reviews program enables you to support fellow scientists by sharing your firsthand experience with our products.
Login to Submit a Review